Fish Pathogen Monitoring Studies
Fish Pathogen Monitoring Studies
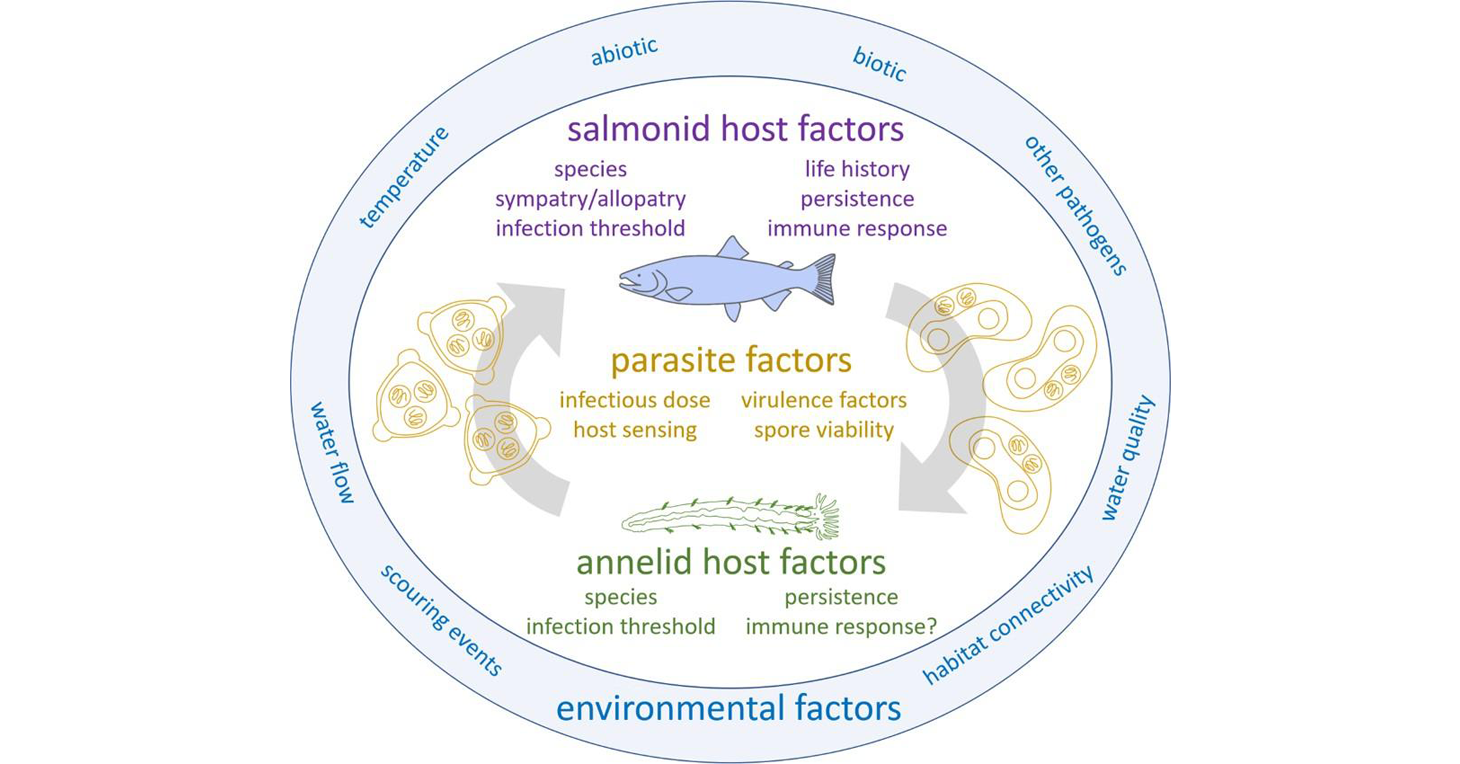
The life cycle of Ceratonova shasta and abiotic and biotic factors that influence the different stages. Published in: Bartholomew JL, Alexander JD, Hallett SL, Alama-Bermejo G, Atkinson SD (2022). Ceratonova shasta: a cnidarian parasite of annelids and salmonids. Parasitology 149, 1862–1875. https://doi.org/10.1017/S0031182022001275
Ceratonova shasta is a freshwater, myxozoan parasite that is native to the Pacific North West of North America. It causes enteronecrosis in juvenile salmonids and is associated with population-level impacts in the Klamath River. Transmission occurs through waterborne stages: actinospores released from annelids infect salmonid fishes and develop into myxospores which then infect annelids (see life cycle, above). The parasite proliferates in each host.
In response to the high prevalence and severity of C. shasta-infection in Klamath salmonids, we developed a parasite monitoring program to track the spatial and temporal abundance of C. shasta. The three main approaches are based on the parasite's life cycle and include sentinel fish exposures, annelid host sampling and molecular quantification of parasite DNA in water samples. These are described in more detail below. Monitoring occurs at established index sites which are shown on the following map.
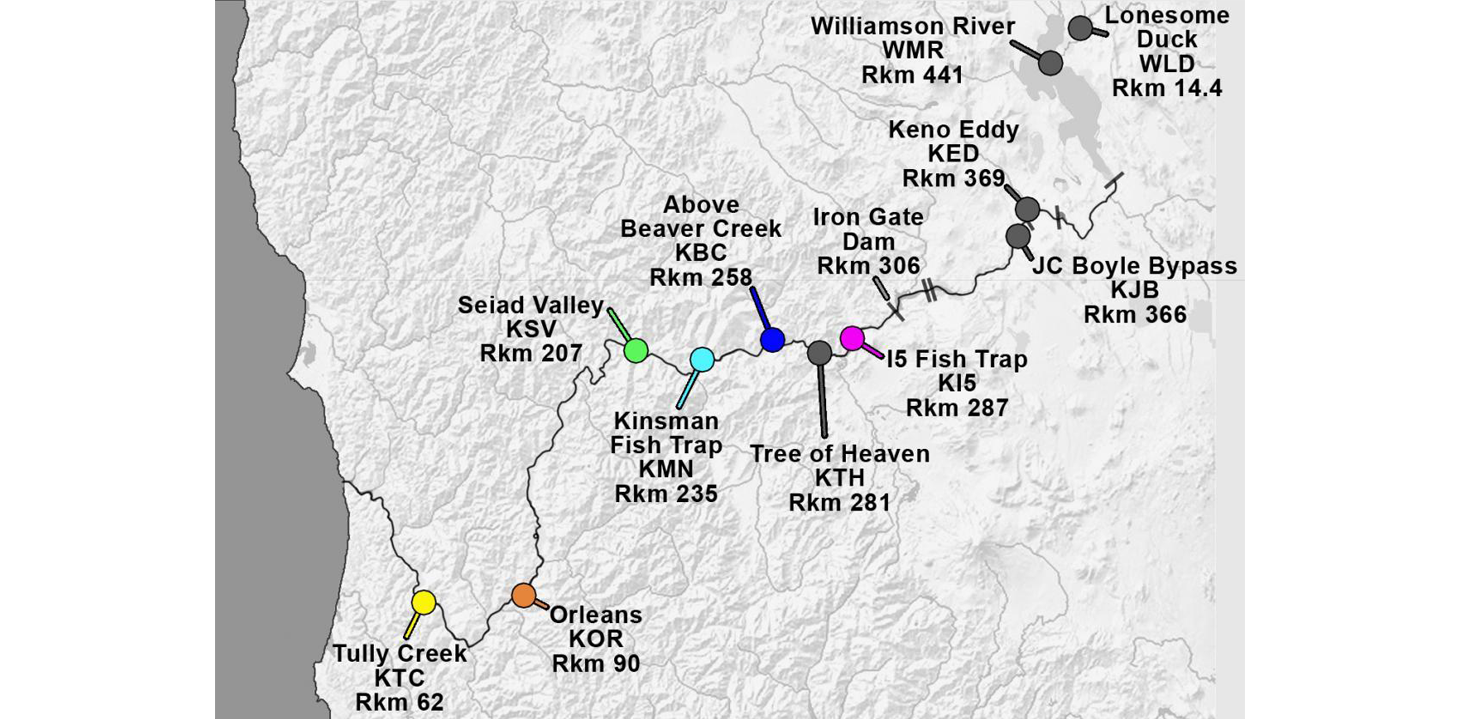
Klamath River Index Sites with site abbreviations and river kilometers (Rkm). Iron Gate Dam (Rkm 306) blocks anadromous salmonid migration.
Data shared here are preliminary and subject to modification.
Page photo credits: S Atkinson, S Hallett & J Alexander
Monitoring Studies are Primarily Supported by the Bureau of Reclamation.
Sentinel fish exposures
In sentinel fish exposures, fish highly susceptible to the parasite (out-of-basin rainbow trout) are placed in cages alongside fish of interest such as in-basin Chinook and coho salmon at index sites along the river for a three day exposure. All fishes are transported to OSU's John L Fryer Aquatic Animal Health Lab and monitored for infection (~ 60 days). Severity of infection (percent morbidity and mean days to morbidity) are recorded through visual observations and molecular assay (PCR).
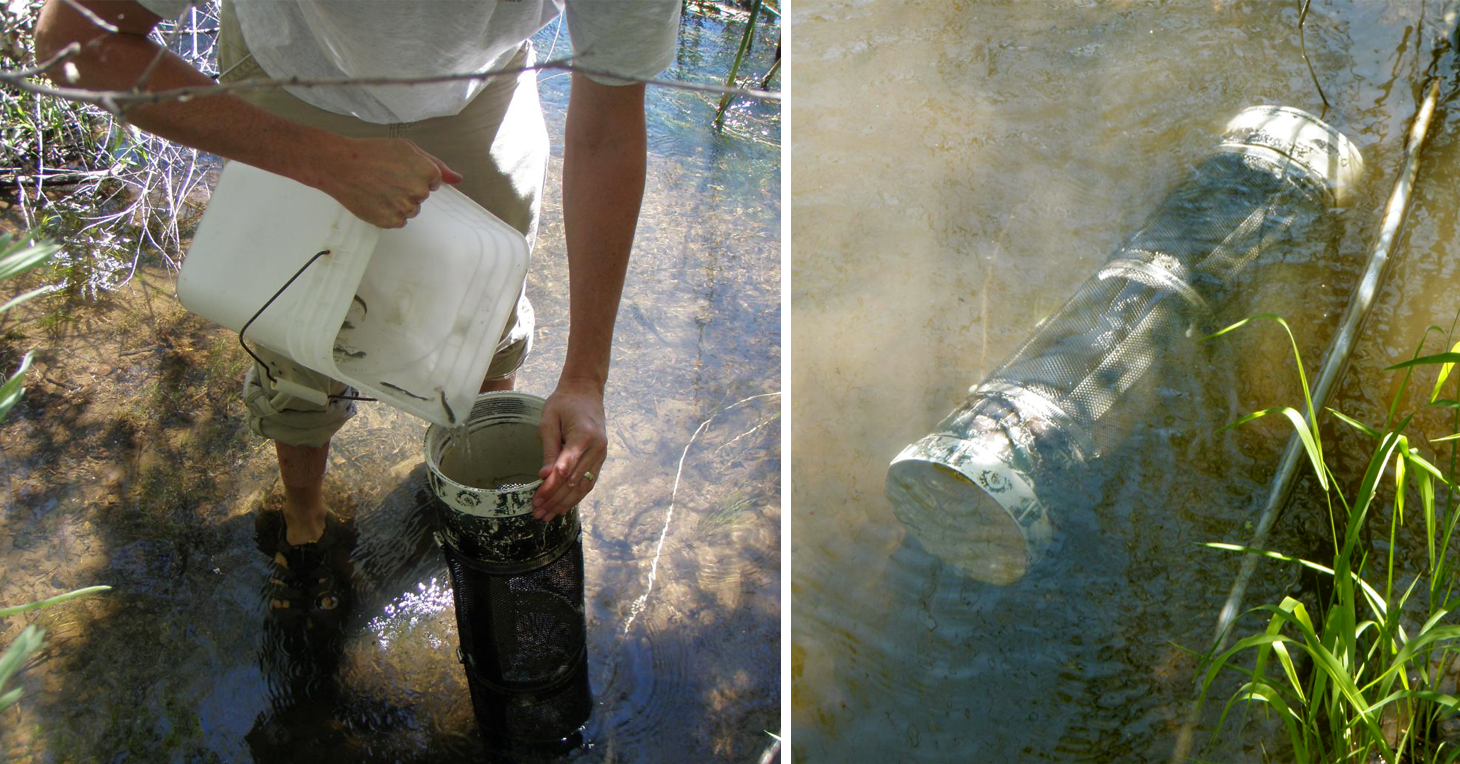

Juvenile fish being placed in a sentinel cage which is then submerged in the Klamath River
2023 data updates
An April sentinel fish exposure was scheduled for 04/19 - 04/22 at KBC and KSV with IGH Fall Chinook and out-of-basin rainbow trout. However, this was postponed to avoid overlapping with the peak of the surface flushing flow event that commenced April 19 and will now occur April 27 - 30.
View a summary of the April sentinel fish exposures here (pdf).
A May sentinel fish exposure occurred 05/25 - 05/28 at WMR and KED in the upper basin and at KI5, KBC, KSV and KOR in the lower basin.
View a summary of the May sentinel fish exposures (pdf).
A third sentinel fish exposure occurred June 14–17, 2023.
View a summary of the June sentinel fish exposures here (pdf).
An exposure occurred from the 19th-22nd September at two lower basin sites.
Water samples
To detect and quantify waterborne stages of C. shasta, river water samples are collected at five mainstem index sites; once a week all year round at two sites (KBC and KSV) and once per week from April through October at three other sites. Solar-powered automatic samplers (ISCOs) collect 1L water every 2 hours for 24 hours, from which 4 1L samples are manually taken (see photos below). Each 1L sample is filtered through a nitrocellulose membrane using a vacuum pump, and any captured DNA in 3 of the replicate samples is extracted using a kit (see images on right). A quantitative PCR (qPCR) specific for C. shasta is used to detect and quantify any parasite DNA present. Cq values generated by the qPCR are converted to numbers of parasite spores per liter of water using reference samples with known quantities of spores. The Karuk and Yurok tribal biologists are integral to the collection and filtration of the water samples. Water samples are also taken in conjunction with the sentinel fish exposures; manual 'grab' samples are collected on the first and last day of the exposure. Data are presented as the average spores per liter of three replicate 1 liter water samples collected at each site and time.
Genotyping
There are multiple genetic types or genotypes of C. shasta simultaneously present in the Klamath River. These differentially disease the various salmonid species. For example, type I causes mortality in Chinook salmon whereas type II can be fatal for coho salmon. Type 0 is found in sympatric Oncorhynchus mykiss (steelhead and redband rainbow trout).
Therefore, we also genotype each water sample. This is done in two ways. For genotyping type II only, we use a qPCR that amplifies the variable ITS1 region specific to that genotype. Whereas, to determine the overall genotype composition of a sample, we amplify the variable ITS1 region common to all genotypes in a PCR and then sequence that amplicon. From the sequencing chromatogram, we can determine the proportion of each genotype present in a sample. We use the total spore density to then determine the number of spores of each genotype in a sample.
Genotyping commences once we detect greater than 1 spore per liter.
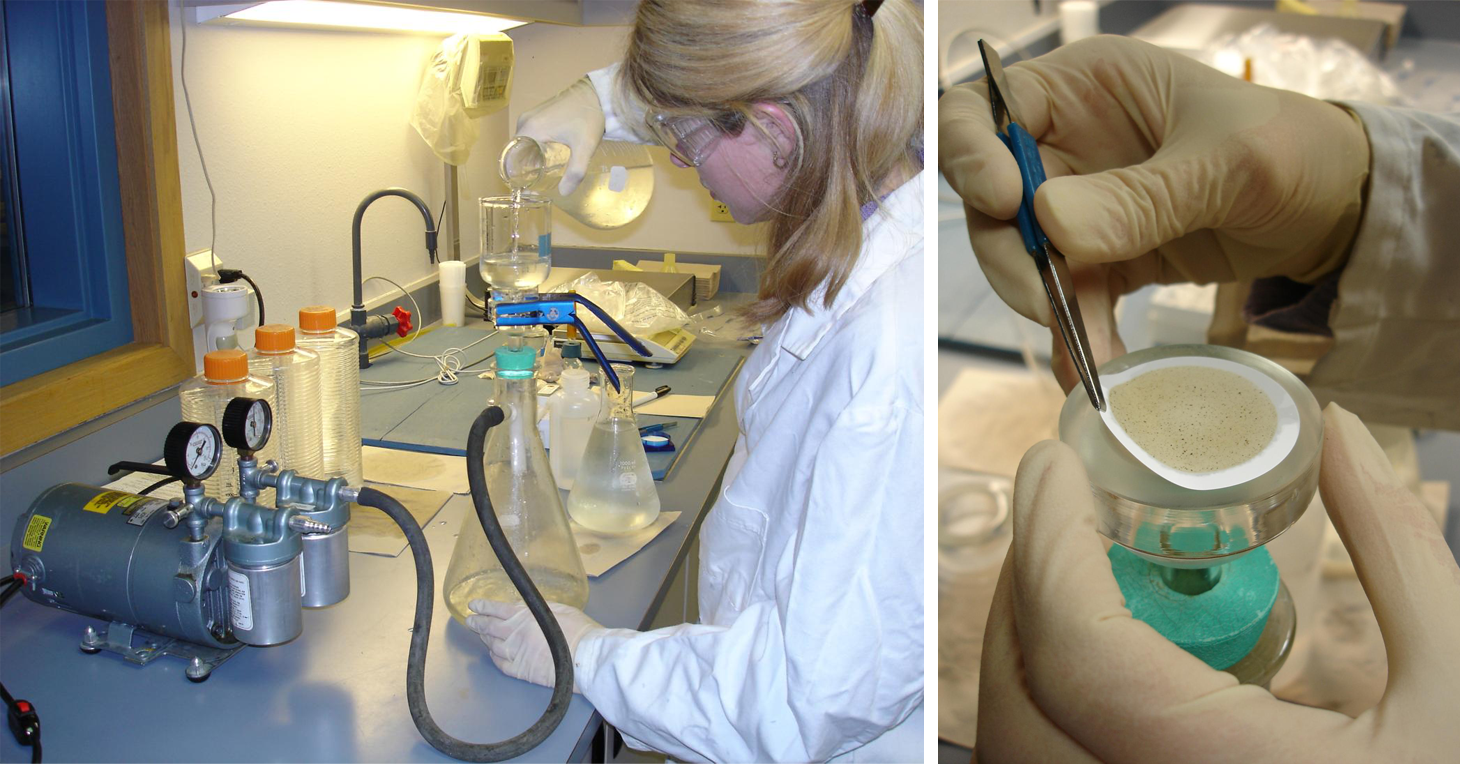

Filtering a water sample using a vacuum pump (left). Folding the filter paper with captured material (right).
2023 data updates
We are still in the Winter collection phase, with only two Klamath River sites sampled each week: KSV & KBC. OSU has received and analyzed water samples up to and including 2/13/2023.
Samples were taken either by ISCO automatic sampler "i" and are given as the average density of C. shasta spores per liter of river water (from 3x 1-L subsamples of a 24-hour composite sample, Sunday-Monday), or taken manually "g" (3 x 1-L samples taken Monday).
Samples from late January and into February have had fewer contaminants (sediment, organics) meaning less assay inhibition, although a low level of inhibition persists in most samples - likely due to continued runoff from fire damaged areas.
No C. shasta DNA has been detected in water since trace detections over the New Year. These very low to no detections of C. shasta DNA are typical of early-mid winter samples every year.
Specific results for 2 sites, spores/L, and indicating whether the sample was taken by automatic ISCO sampler (i) or manual grab (g).
| KBC | KSV | |
| 02/13/2023 | i 0 | i 0 |
| 02/06/2023 | g 0 | g 0 |
| 01/30/2023 | g 0 | i 0 |
| 01/23/2023 | g 0 | g 0 |
| 01/17/2023 | i 0 | g 0 |
No C. shasta DNA was detected in samples collected at KBC and KSV.
Sampling at all index sites will commence late March/early April.
Winter season sampling continues with 2 sites sampled each week: KBC & KSV. No C. shasta DNA was detected in water samples taken up to 3/20/2023, the most recent samples received and analyzed by OSU.
This was the final week of the regular restricted winter sampling: at only 2 index sites. Both sites were sampled by automatic ("ISCO"; i) sampler. Site KBC required twice the usual number of filters due to high levels of suspended material (presumably residual burn-related sediment being mobilized by ongoing rain events); this material also causes inhibition of the DNA assay which can interfere with detection.
KBC i, no Cs-DNA detected, moderate inhibition
KSV i, no Cs-DNA detected, no inhibition
This was the first week of sampling at all 6 index sites, with expedited sample processing. Multiple sites required double the regular number of filters due to high levels of suspended material. Four sites were sampled with an automatic sampler ("i"), site KOR was a manual grab ("g").
No Cs-DNA was detected at any of the 6 sites analyzed:
KI5 i, no Cs-DNA detected, slight inhibition of 1/3 samples
KBC i, no Cs-DNA detected, but high inhibition of all samples
KMN i, no Cs-DNA detected, slight inhibition of 2/3 samples
KSV i, no Cs-DNA detected, no inhibition
KOR g, no Cs-DNA detected, no inhibition
KTC g, no Cs-DNA detected, no inhibition
4/10/2023 was the 2nd week of sampling at all 6 index sites, with expedited sample processing. Multiple sites required double the regular number of filters due to high levels of suspended material (presumably residual burn-related sediment being mobilized by ongoing rain events); this material also causes inhibition of the DNA assay which can interfere with detection. Two sites were sampled with automatic sampler ("i"), the rest were manual grabs ("g").
No Cs-DNA was detected at any of the 6 sites analyzed:
KI5 i, no Cs-DNA detected, no inhibition
KBC i, no Cs-DNA detected, no inhibition
KMN g, no Cs-DNA detected, inhibition in 1/3 samples
KSV g, no Cs-DNA detected, inhibition in 2/3 samples
KOR g, no Cs-DNA detected, slight inhibition in 2/3 samples
KTC g, no Cs-DNA detected, no inhibition
4/17/2023 was the 3nd week of sampling at all 6 index sites, with expedited sample processing.
For the first time this year, C.shasta-DNA was detected in water samples - at levels less than 1 spore per liter, and at 4 sites.
3 sites were sampled with automatic sampler ("i"), 3 were manual grabs ("g").
Again, multiple sites required double the regular number of filters due to high levels of suspended material (presumably residual burn-related sediment being mobilized by ongoing rain events); this material also causes inhibition of the DNA assay which can interfere with detection.
The density of C. shasta at each site is as follows:
KI5 i, no Cs-DNA detected, low inhibition in all samples
KBC i, <1 spore/liter (initial test fully inhibited, sample repurified and re-assayed)
KMN g, <1 spore/liter, low inhibition in 1/3 samples
KSV i, <1 spore/liter, low inhibition in 2/3 samples
KOR g, <1 sp/liter, no inhibition
KTC g, no Cs-DNA detected, no inhibition
4/24/2023 was the 4th week of sampling at all 6 index sites, with expedited sample processing. This week's sampling occurred during the Surface Flushing Flow event.
Despite higher levels of contaminents# (likely due to the flushing flow), for the first time this year, C. shasta DNA was detectable at all sites.
3 sites were sampled with automatic sampler ("i"), 3 were manual grabs ("g").
Most sites required double the regular number of filters due to #high levels of suspended material (likely due to the flushing flow); this material also causes inhibition of the DNA assay which can interfere with detection.
In anticipation of inhibition in some samples, we ran samples in duplicate with one set at a calibrated higher dilution (indicated by a * below) and these calibrated values are reported when the undiluted regular samples were inhibited.
The density of C. shasta at each site is as follows:
KI5 i, <1 spore/liter; 2/3 samples clean, 1/3 low inhibition
KBC i, <1 sp/L* (undiluted DNA was fully inhibited)
KMN g, 3 sp/L* (undiluted DNA was fully inhibited)
KSV g, 1 sp/L*, (undiluted DNA was fully inhibited)
KOR i, <1 sp/L, weak inhibition in 2/3 samples
KTC g, <1 sp/L, no inhibition
05/01/2023 was the 5th week of sampling at all 6 index sites, with expedited sample processing. This sampling occurred immediately after the Surface Flushing Flow event.
All sites were sampled with automatic sampler ("i"), except KTC, which was a manual grab ("g").
Samples from KBC downstream continue to have sediment and suspended organic material in them. Thus most sites required double the regular number of filters due to the high levels of suspended material (likely due to the residual flushing flow and ongoing runoff). This material also causes inhibition of the DNA assay which can interfere with detection of target DNA.
In anticipation of inhibition in some samples, we ran samples in duplicate with one set at a calibrated higher dilution (indicated by a * below) and these calibrated values are reported when the undiluted regular samples were inhibited. Samples reported here as inhibited will be rerun and reported next week.
C. shasta-DNA was detectable at all sites where a reading could be obtained. The density of C. shasta at each site is as follows:
KI5 i, 2 spores/liter; no inhibition
KBC i, 7 sp/L* (undiluted DNA was highly inhibited)
KMN i, 7 sp/L* (undiluted DNA was fully inhibited)
KSV i, 3 sp/L* (undiluted DNA was fully inhibited)
KOR i, 4 sp/L* (undiluted DNA was fully inhibited)
KTC g, 1 sp/L; no inhibition
Samples with at least 5 sp/L will be analysed with the coho assay and reported next week, and from next week we will run the coho assay in addition to total spore assay.
05/08/2023 was the 6th week of sampling at all 6 index sites, with expedited# sample processing. (#Sample shipment was delayed 1 business day due to courier).
All sites were sampled with automatic sampler ("i"), except KTC and KOR, which were a manual grab ("g").
Samples from KBC downstream continue to have sediment and suspended organic material in them, which necessitates double the regular number of filters and causes inhibition of the DNA assay which can interfere with detection. C. shasta-DNA was detectable at all sites where a reading could be obtained, and at higher levels than previous weeks (1-10 fold increase, with 10 or more spores per liter measured). The results from the second assay (to detect 'coho' Cs genotype) are in progress.
Samples that we anticipated from previous weeks would have inhibition, we ran at a higher dilution or reprocessed (indicated by a * below). Some samples had residual inhibition effects and will be rerun and reported with next week's samples (including KBC which remained inhibited even after dilution and therefore requires further processing).
The density of C. shasta at each site is as follows:
KI5 i, 13 spores/liter; no inhibition
KBC i, 13 sp/L* (DNA was highly inhibited - was reprocessed); <1 sp/L II-Coho
KMN i, 22 sp/L* (undiluted DNA was fully inhibited)
KSV i, 12 sp/L* (undiluted DNA was fully inhibited)
KOR i, 11 sp/L* (DNA was highly inhibited - was reprocessed); <1 sp/L II-Coho
KTC g, 10 sp/L*; (undiluted DNA was inhibited)
5/15/2023 was the 7th week of sampling at all 6 index sites, with expedited sample processing.
Only two sites were sampled with automatic sampler ("i" - KI5, KBC), the rest were by manual grab ("g").
All sites required our tribal collaborators to use 2 filters instead of 1 per liter, indicating ongoing high levels of sediment or organic debris. As this material also causes inhibition of the DNA assay, which can interfere with detection (and we have seen evidence of this most weeks this year), we preemptively ran all of this week's sites at a higher dilution to reduce potential inhibition; this was successful. C. shasta-DNA was detectable at all sites, but at lower levels than last week.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at all sites, at a level of 10-38% of the total C. shasta present.
The density of C. shasta at each site is as follows:
KI5 i, 4 (<1 coho) spores/liter
KBC i, 2 (<1 coho) sp/L
KMN g, 8 (3 coho) sp/L
KSV g, 5 (1 coho) sp/L
KOR g, 5 (<1 coho) sp/L
KTC g, 4 (<1 coho) sp/L
5/22/2023 was the 8th week of sampling at all 6 index sites, with expedited sample processing.
Only two sites were sampled by manual grab ("g" KMN, KOR), the rest were with automatic sampler ("i"). KTC was sampled both ways.
All sites required our tribal collaborators to use 2 filters instead of 1 per liter, indicating ongoing high levels of sediment or organic debris. As this material also causes inhibition of the DNA assay, which can interfere with detection (and we have seen evidence of this most weeks this year), we preemptively ran all of this week's sites at a higher dilution to reduce potential inhibition; this was successful. C. shasta-DNA was detectable at all sites, some at higher levels and some at lower levels than last week.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at all sites.
The density of C. shasta at each site is as follows:
KI5 i, 6 (2 coho) spores/liter
KBC i, 6 (1 coho) sp/L
KMN g, 13 (4 coho) sp/L
KSV i, 6 (2 coho) sp/L
KOR g, 6 (2 coho) sp/L
KTC g, 4 (1 coho) sp/L
KTC i, 3 (<1 coho) sp/L
05/29/2023 was the 9th week of sampling at all 6 index sites, with expedited sample processing.
Three sites were sampled by manual grab ("g", KI5, KSV, KOR), the other three were with automatic sampler ("i").
All sites except KI5 and KOR required our tribal collaborators to use 2 filters instead of 1 per liter, indicating ongoing high levels of sediment or organic debris. As this material also causes inhibition of the DNA assay, which can interfere with detection (and we have seen evidence of this most weeks this year) we preemptively ran all of this week's sites at a higher dilution to reduce potential inhibition; this was successful. C. shasta-DNA was detectable at all sites.
The three samples analyzed from site KSV had high standard deviation between them, so the reported value should be regarded as only indicative of the spore level at this point - we will process the reserved 4th sample and re-analyze the data.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses.
The density of C. shasta at each site is as follows:
KI5 g, 19 (15 coho) spores/liter
KBC i, 10 (7 coho) sp/L
KMN i, 13 (6 coho) sp/L
KSV g, 6 (4 coho) sp/L (high standard deviation between samples))
KOR g, 6 (4 coho) sp/L
KTC i, 3 (2 coho) sp/L
Observations:
C. shasta-DNA, including type II, was detected at all index sites this week.
Spore densities were either similar or higher this week compared to the previous week. They were higher at the uppermost three sites and lowest at the lowermost three sites.
Total spore densities exceeded 10 spores per liter at three index sites and coho type II densities were above 5 spores per liter at those same three sites (uppermost sites). The proportion of type II in a sample was as high as ¾.
06/05/2023 was the 10th week of sampling at all 6 index sites, with expedited sample processing.
All sites except KOR were sampled by automatic sampler ("i").
Water quality (suspended particles) has improved at the lowermost 2 sites (KOR & KTC), which only required a single filter for each liter, however the other sites still required 2 filters per liter, and KBC in particular contained a lot of sediment. As this material also causes inhibition of the DNA assay, which can interfere with detection (and we have seen evidence of this most weeks this year) we again preemptively ran all of this week's sites at a higher dilution to reduce potential inhibition; this was successful. C. shasta DNA was detectable at all sites.
A single sample from each of KSV and KBC ran as significant outliers to the replicate bottles, so the usually reserved 4th sample was processed. We did this also for KSV last week (which had the same issue) and have now updated the graph with the corrected value.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at all sites.
Total spore densities were considerably lower at he two upermost index sites and were above 10 spores per liter only at one site this week (Kinsman - KMN) and coho type II density was also above 5 spores per liter only at this site.
The density of C. shasta at each site is as follows:
KI5 i, 3 (1 coho) spores/liter
KBC i, <1 (<1 coho) sp/L (high standard deviation among samples; n=4)
KMN i, 14 (6 coho) sp/L
KSV i, 4 (2 coho) sp/L (high standard deviation among samples; n=4)
KOR g, 4 (1 coho) sp/L
KTC i, 4 (1 coho) sp/L
The vertical scale of the all-sites graph has been adjusted to match the KBC graph, to give better resolution of all points.
6/12/2023 was the 11th week of sampling at all 6 index sites, and starting this coming week analyses will no longer be expedited as we are out of the peak outmigrant season.
Debris flows in the basin significantly increased turbidity and both field and lab crews struggled with filtering and purifying the dirtier water. Only one sample from KI5 was filtered on one filter membrane, all other samples & sites required at least 2 or 4 filters per liter. As this material also causes inhibition of the DNA assay, which can interfere with detection (and we have seen evidence of this most weeks this year) we again preemptively ran all of this week's sites at a higher dilution to reduce potential inhibition; this was successful. C. shasta DNA was detectable at all sites.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at low levels at all sites except KOR (negative).
The density of C. shasta at each site is as follows:
KI5 i, 5 (3 coho) spores/liter
KBC g, 3 (2 coho) sp/L
KMN g, 1 (<1 coho) sp/L
KSV i, 1 (<1 coho) sp/L (high standard deviation among samples)
KOR g, <1 (0 coho) sp/L
KTC i, 1 (<1 coho) sp/L
6/19/2023 was a holiday so samples that week were collected Tuesday.
6/20/2023 was the 12th and final week of sampling at all 6 index sites.
River water filtered much easier this week as evidenced by tribal collaborators only needing 1 filter per sample at most sites.
Spore densities exceded 10 spores per liter at every index site and were dramatically higher at the uppermost 2 or three sites, having the highest values seen in the lower basin so far this season.
As spore levels were expected to be >5sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at all sites, with > 5 spores per liter detected at half of these.
The density of C. shasta at each site is as follows:
KI5 i, 67 (24 coho) spores/liter
KBC i, 22 (3 coho) sp/L
KMN g, 43 (11 coho) sp/L - final sample for the season at this site
KSV i, 15 (5 coho) sp/L
KOR g, 11 (3 coho) sp/L
KTC i, 12 (2 coho) sp/L
*Updated graphs will be forthcoming*
6/26/2023 was the first week of sampling at 5 index sites (excluding KMN).
River water filtered more easily again this week as evidenced by tribal collaborators only needing 1 filter per sample at most sites.
For both this and last week, spore densities are dramatically higher at the uppermost 2 or three sites, having the highest values seen in the lower basin so far this season.
As spore levels were expected to be >5 sp/L, we ran a second assay to determine the amount of C. shasta genotype II present (the type that infects Coho). Those numbers are reported in parentheses. This "Coho genotype" was detected at all sites, exceding 5 spores per liter at 2 of these.
The density of C. shasta at each site 6/26/2023 is as follows:
KI5 g, 61 (24 coho) spores/liter
KBC g, 77 (6 coho) sp/L
KSV g, 14 (2 coho) sp/L
KOR g, 3 (<1 coho) sp/L
KTC g, 10 (1 coho) sp/L
7/3/2023 was a holiday week so samples that week were collected Wednesday 7/5.
River water filtered more easily this week, as evidenced by tribal collaborators only needing 1 filter per sample at most sites; however, site KSV continues to have intermittent high sediment and inhibition.
Spore densities remained some of the highest seen this year, with a shift in higher values downstream this week.
Coho genotyping is finished for the year.
Total C. shasta spores per liter this week are as follows:
KI5 i, 9 spores/liter
KBC g, 73 sp/L
KSV i, 15 sp/L
KOR g, 54 sp/L
KTC i, 22 sp/L (KTC g, 44 sp/L)
Spore abundance was lower this week at all sites enumerated this week except KSV, but still remained above 10 spores per liter at 4 of the 5 sites (same as last week).
The density of C. shasta at each site 7/10/2023 was as follows:
KI5 i, 4 sp/L
KBC g, 43 sp/L
KSV g, 40 sp/L
KOR i, 52 sp/L
KTC i, 33 sp/L
River water clarity has remained good - as evidenced by tribal collaborators only needing 1 filter per sample at most sites. The density of waterborne C. shasta was lower this week at all sites. Densities were highest at the lowermost sites with 10 or more spores per liter measured at 3 of the 5 sites.
The density of C. shasta at each site 7/17/2023 was as follows:
KI5 i, 2 spores/liter
KBC i, <1 sp/L
KSV g, 16 sp/L
KOR i, 10 sp/L
KTC i, 23 sp/L
River water clarity has remained good - as evidenced by tribal collaborators only needing 1 filter per sample at most sites. The density of waterborne C. shasta was highest at the two lowermost sites (KOR and KTC) and only at these were more than 10 spores per liter measured.
The density of C. shasta at each site 7/24/2023 was as follows:
KI5 g, 7 sp/L
KBC i, 5 sp/L
KSV g, 6 sp/L
KOR g, 17 sp/L
KTC i, 18 sp/L
River water clarity was good on 7/31 as evidenced by tribal collaborators only needing 1 filter per sample at most sites (notable exception is KOR which had a lot of sediment).
The density of waterborne C. shasta was highest at KSV and more than 10 spores per liter were measured at two of the five sites.
7/31/2023
KI5 g, 3 spores/liter
KBC i, 0 sp/L
KSV g, 23 sp/L
KOR i, 2 sp/L
KTC i, 18 sp/L
River water clarity was worse this week with most sites needing double the amount of filters.
The density of waterborne C. shasta was highest again this week at KSV and more than 10 spores per liter were measured at three of the five sites (the three lowermost sites).
8/07/2023
KI5 g, 5 sp/L
KBC i, 4 sp/L
KSV g, 42 sp/L
KOR g, 29 sp/L
KTC i, 14 sp/L
The density of waterborne C. shasta was once again highest at KSV (third consecutive week) and more than 10 spores per liter were measured at two of the five sites.
KI5 g, 4 spores/liter
KBC i, 3 sp/L
KSV g, 23 sp/L
KOR i, <1 sp/L
KTC i, 14 sp/L
Wildfires prevented water sampling this week at all sites except KTC:
KI5 x
KBC x
KSV x
KOR x
KTC i, 5 sp/L
Water quality was markedly poorer this week, as evidenced by multiple water filters being needed at all sites. KBC was unable to be sampled. More than 10 spores per liter were measured at two of the five sites.
KI5 g, 13 spores/liter
KBC x
KSV g, 7 sp/L
KOR i, not detected
KTC i, 12 sp/L
Water quality remained poor, as evidenced by multiple water filters required at all sites. KTC samples will be shipped to OSU next time. Fewer than 10 spores per liter were detected at all index sites measured (first time since June 12th).
KI5 g, 8 spores/liter
KBC g, 4 sp/L
KSV g, 8 sp/L
KOR g, <1 sp/L
KTC TBD
The density of waterborne C. shasta remained below 10 spores per liter and decreased further at all index sites this week:
KI5 g, 1 spores/liter
KBC i, inhibited - will be rerun
KSV g, <1 sp/L
KOR i, 2 sp/L
KTC g, 3 sp/L
Waterborne C. shasta was detected at all sites in the lower basin this week, however, densities were low:
KI5 g, 1 sp/L
KBC i, inhibited - will be rerun
KSV g, 1 sp/L
KOR i, 1 sp/L
KTC g, 1 sp/L
Low densities of waterborne C. shasta were detectable at all sites where samples were taken. No data are available yet for KTC. The KBC graph has been updated, but the reruns of inhibited samples from prior weeks have not yet been included.
KI5 g, 2 spores/liter
KBC i, 2 sp/L
KSV i, 1 sp/L
KOR i, 2 sp/L
KTC n/d
Low densities of waterborne C. shasta were detectable at all sites where samples were collected. Theft of the I5 sampler meant no samples there 9/30. No data are available yet for KOR 9/30 and KTC. The KBC graph has been updated, but does not include the reruns of inhibited samples from prior weeks.
KI5 n/d [water sampler stolen]
KBC i, 2 sp/L
KSV i, 1 sp/L
KOR n/d
KTC n/d
Waterborne C. shasta was detectable at all sites where samples were taken. Low densities were measured throughout the lower basin.
KI5 g, 2 spores/liter
KBC i, <1 sp/L
KSV i, <1 sp/L
KOR i, <1 sp/L
KTC g, 5 sp/L
Waterborne C. shasta was detectable at all sites where samples were taken (except KBC - no data). Low densities were measured throughout the lower basin. KBC (and to some extent KSV) samples had high levels of sediment, rendering them difficult to filter, and difficult to process for analysis.
KI5 g, 3 sp/L
KBC i, no data - high sediment
KSV i, 1 sp/L
KOR i, <1 sp/L
KTC g, 1 sp/L
Waterborne C. shasta was detectable at all sites where samples were taken (except KBC - no data). Low densities were measured throughout the lower basin. KBC (and to some extent KSV) samples had high levels of sediment, rendering them difficult to filter, and difficult to process for analysis.
KI5 g, 1 sp/L
KBC i, no data - high sediment
KSV i, trace sp/L
KOR i, <1 sp/L
KTC g, 2 sp/L
As is typical in Fall, the densities of C. shasta are low. The parasite was still detected throughout the lower basin but generally just a few spores per liter (the exception was KBC with 7 spores per liter measured, but the sample differed this week in that it had a high sediment load).
KI5 1 g
KBC 7 g (was rerun due to inhibition; plentiful sediment may affect the sample)
KSV <1 i
KOR <1 i
KTC 2 g
Low quantities of waterborne C. shasta were detected throughout the lower basin.
KI5 2 g
KBC 1 g (was rerun due to inhibition; plentiful sediment in the sample)
KSV 3 g
KOR 1 i
KTC 1 g
Water sampling is occurring at only two index sites from November through March: KBC and KSV. Low quantities of waterborne C. shasta (< 1 spore per liter) were detected from November onwards.
Density (average spores per liter) of Ceratonova shasta in 24-hour composite water samples collected at the mainstem long term index site, near Beaver Creek (KBC) in 2023 (blue data points) compared to the previous 5 years (2018 - 2022). A managed surface flushing flow event commenced April 19th (6030 cfs for 3 days).
Density (average spores per liter) of Ceratonova shasta in 24-hour composite water samples collected at the mainstem index sites in 2023. Note that KMN is sampled only during salmonid outmigration, KBC and KSV year round and remaining sites April through October. KBC = near Beaver Creek, KSV = Seiad Valley, KI5 = near I5 bridge, KTC = Tully Creek, KMN = Kinsman Fish Trap, KOR = Orleans. The line denotes 10 spores per liter which corresponds with 40% mortality threshold in Chinook salmon. A managed surface flushing flow event commenced April 19th (6030 cfs for 3 days).
Density (average spores per liter) of Ceratonova shasta in 24-hour composite water samples collected at the mainstem index sites in 2022. Note that KMN is sampled only during salmonid outmigration, KBC and KSV year round and remaining sites April through October. KBC = near Beaver Creek, KSV = Seiad Valley, KI5 = near I5 bridge, KTC = Tully Creek, KMN = Kinsman Fish Trap, KOR = Orleans. The line denotes 10 spores per liter which corresponds with 40% mortality threshold in Chinook salmon.
Density (average spores per liter) of Ceratonova shasta genotype II in 24-hour composite water samples collected at the mainstem index sites in 2021. Note that KMN is sampled only during salmonid outmigration, KBC and KSV year round and remaining sites April through October. KBC = near Beaver Creek, KSV = Seiad Valley, KI5 = near I5 bridge, KTC = Tully Creek, KMN = Kinsman Fish Trap, KOR = Orleans. The line denotes 5 spores per liter which corresponds with a 40% mortality threhold in coho salmon.
Manual water sampling by Karuk Tribal Biologist Larry Alameda at index site K-I5
Solar powered automatic sampler at the Kinsman index site. An ISCO is used at all index sites but manual sampling provides a backup.
Collection of 4 1L technical replicate water samples from the larger 12L composite collected over 24 hours at KSV
Manual water sampling at Orleans index site.
Photos courtesy of Karuk Tribal Biologist Larry Alameda
Annelid sampling
To monitor abundance and prevalence of C. shasta infection in the invertebrate annelid host, Manayunkia occidentalis, benthic samples are collected annually at seven sites in the Klamath River. Sites span a discharge gradient; 2 are located in the upper basin downstream from Keno Dam, 3 are located in the middle basin downstream from Iron Gate Dam, and 2 are located in the lower basin downstream from the Scott River. Samples are routinely collected in fall, winter, spring, and summer, and are scheduled to occur more frequently if flooding or pulse flow events are planned. Samples are processed to determine density, simple demographics, and prevalence of C. shasta infection.
Manayunkia occidentalis tubes from the image on right
A high density assemblage sample at TOH
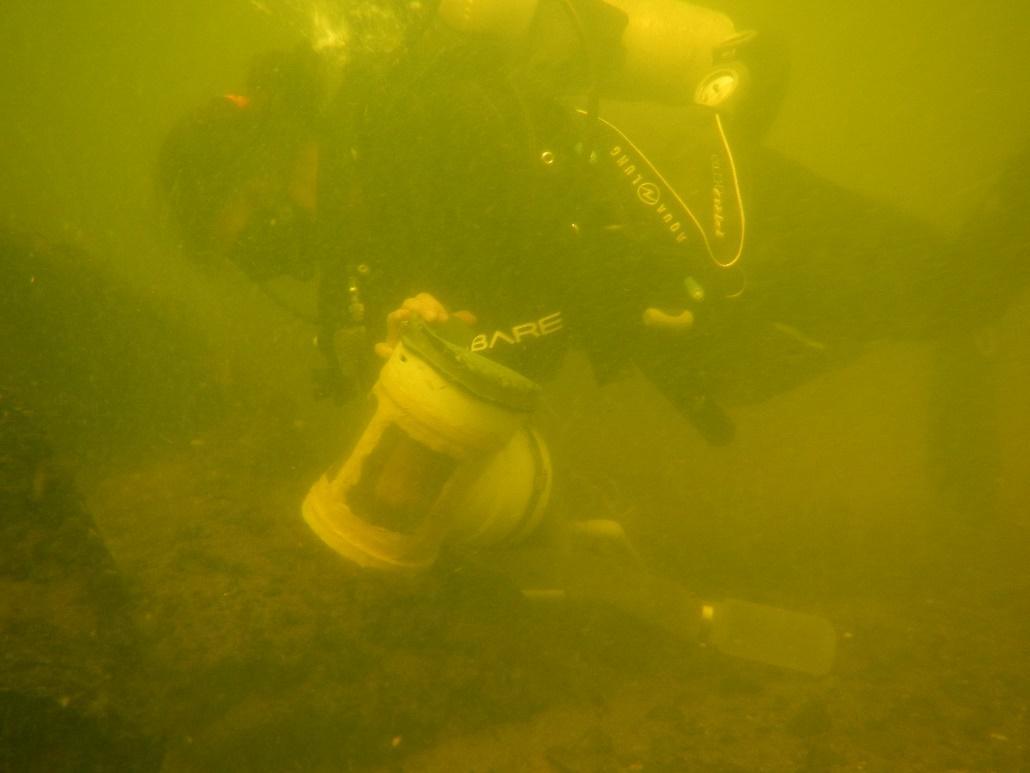

Diver collecting a benthic sample from boulder substrate at the TOH reach. We use a modified Hess sampler fitted with 80um Nytex mesh netting and a 500mL Nalgene collection bottle. Samples are preserved in ethanol until processing.
Benthic samples are fractioned prior to sorting.
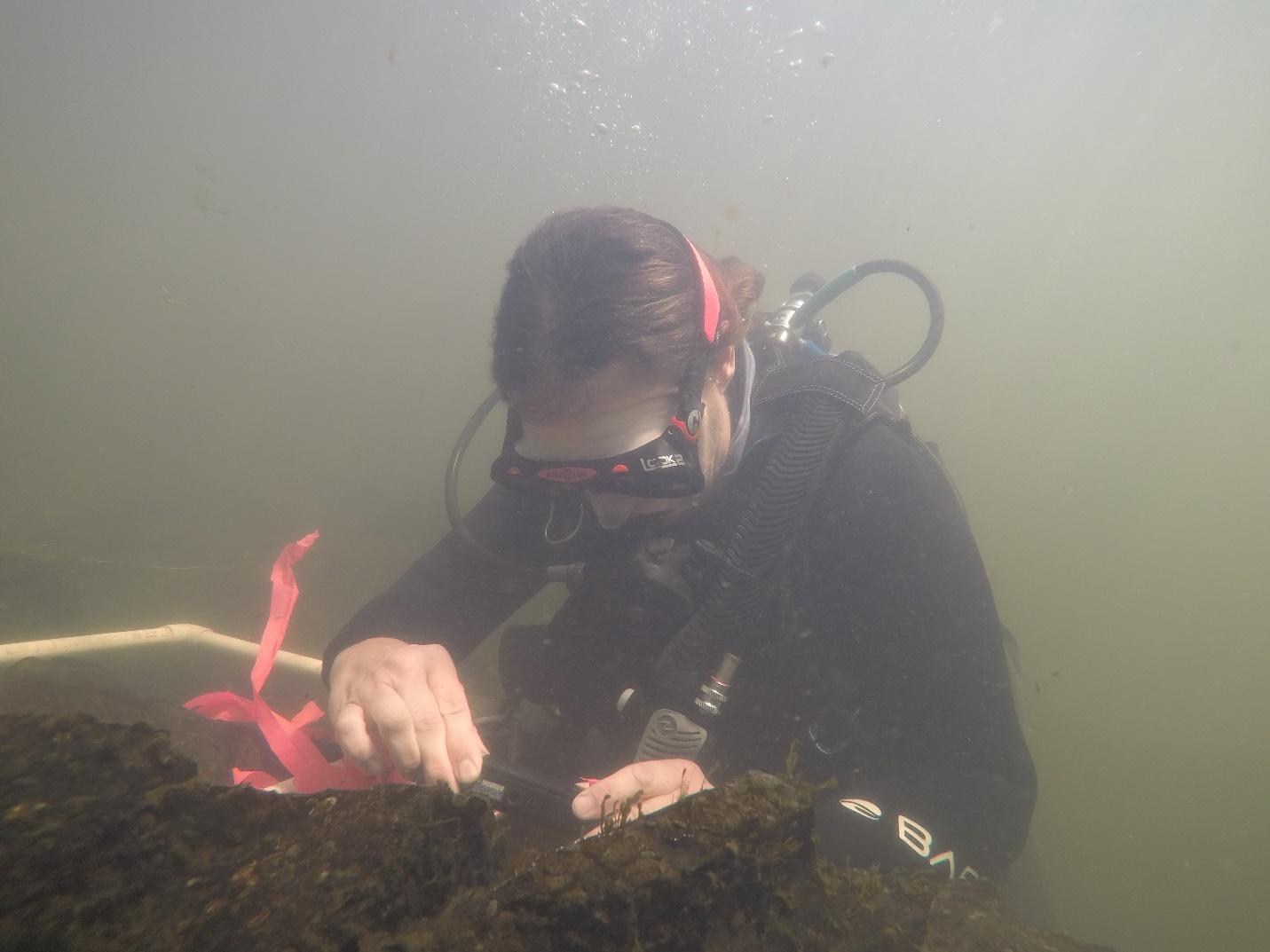

Dr. Alexander photographing annelids on riprap substrate at the KBC monitoring site.
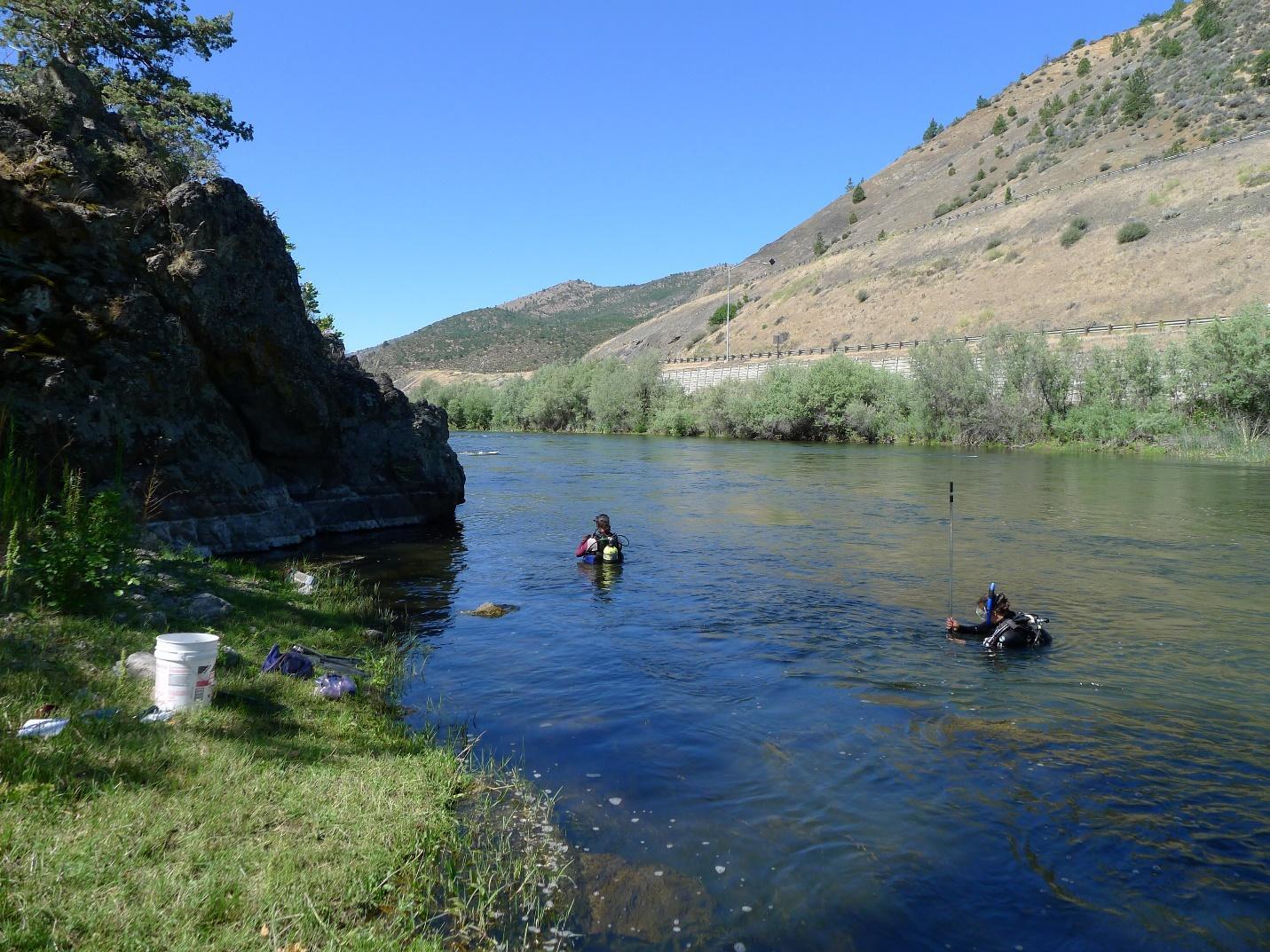

Annelid sampling at the KI5 reach. Foreground diver measures depth and velocity, while background diver measures substrate grain sizes.
Annual reports
Annual reports for Bureau of Reclamation funded studies for 2015 onwards are available. Please contact Sascha Hallett (halletts@oregonstate.edu). Annual reports are submitted June 1 the year after the research year.
Annual Klamath River fish health workshop
We are pleased to announce this year's Annual Klamath River Fish Health Workshop to be held in Yreka, May 17th & 18th. We gratefully acknowledge our Karuk collaborators for hosting us at their Karuk Tribe Housing Authority, Yreka Apsuun Community Center.
This year, the KRFHW will follow the Spring KBMP meeting to be held Tuesday May 16th at the same venue (please see emails from Randy Turner for KBMP specifics).
The general KRFHW schedule will be as follows:
Workshop: Wednesday May 17th (9am-5pm). Full day, public forum during which Klamath River fish health research updates, overviews, and presentations will be shared with an open forum Q & A session. Presentations will be basin-wide. Plans for 2023 and beyond will also be shared. The May 17 workshop will be open to the public to promote an open exchange of information, and attendance for all is free.
Project Planning Session: Thursday May 18th (9am-12pm). Half day, smaller meeting to coordinate research efforts and management needs. The Project planning session is reserved for fish health researchers to discuss 2023 and beyond study plans and to promote project coordination and collaboration. The planning session will include an opportunity for Q & A between fish health researchers and agency/tribal co-managers.
The meeting format will be hybrid, with in-person attendance highly recommended but in the spirit of inclusivity and accessibility, participants will also be able to connect via online (link will be provided following registration).
There is no registration fee for participation, however, we do request that you register your intention to attend on this spreadsheet.
Call for KRFHW presenters: If you would like to present a research update or overview at the Workshop (May 17 public forum), please send us your research abstract (with title, presenter, co-authors, and summary of research) by Friday April 28th.
We look forward to seeing many familiar faces at this annual workshop on fish health related research in the Klamath River Basin!



